All News
October 4, 2024
Integrative structures in PDB-Dev are now available at wwPDB Digital Object Identifier (DOI) landing pages, along with >225,000 experimental structures in the PDB archive. DOI landing pages are available for both released and on-hold entries in PDB-Dev. These pages present basic information about the corresponding integrative structure, offer download of model coordinates and validation files from the PDB archive (https://files.wwpdb.org/pub/pdb_ihm/), and provide a link to the PDB-Dev resource. The PDB-Dev website has been updated to include the newly available links to PDB DOIs on entry pages.
For an example, visit the DOI landing page for a recently-released integrative structure in the PDB archive via PDB DOI: https://doi.org/10.2210/pdb9a8n/pdb.
PDB DOIs issued for each entry are linked from the online versions of papers where PDB IDs are mentioned. Integrative structures are identified on the wwPDB DOI landing pages using the term "Structure Determination Methodology".
Questions or feedback? Contact pdb-dev@mail.wwpdb.org. For more information visit wwpdb.org.
August 14, 2024
Structures determined by integrative and hybrid structure determination methods (IHM) are now available alongside experimental structures in the PDB archive. These structures are deposited to and processed by the PDB-Dev system. Users can access and download IHM structures and associated data from the PDB archive.
Currently, holding files in JSON format, validation reports (summary and full reports) in PDF format, and model files in PDBx/mmCIF format are provided.
- /pdb_ihm/holdings/ (current holdings, released structures last modified dates, unreleased entries)
- /pdb_ihm/data/entries/hash/{PDB_id}/validation_reports/ (summary and full validation reports)
- /pdb_ihm/data/entries/hash/{PDB_id}/structures/ (latest version of model files)
Data may be expanded in the future based on community needs. In the near future, IHM data will also be available via wwPDB DOI landing pages and wwPDB partner websites.
Questions or feedback? Contact deposit-help@mail.wwpdb.org or pdb-dev@mail.wwpdb.org.
August 14, 2024
Integrative structures deposited to PDB-Dev will now be issued 4-character PDB accession codes (e.g., 8ZZ1). Going forward all new depositions will only be issued PDB accession codes and PDB-Dev accession codes (e.g., PDBDEV_00000001) will not be issued.
Existing entries have been remediated to include new PDB accession codes and re-released. Both accession codes, PDB and PDB-Dev (if available), are reported in the database_2 category in IHMCIF files. Additionally, integrative method provenance is captured at _struct.pdbx_structure_determination_methodology. Validation reports have been updated based on the PDB accession codes.
The PDB-Dev website will support search and access of integrative structures using both PDB and PDB-Dev accession codes.
August 14, 2024
IHMCIF, the data standard underpinning the PDB-Dev system, has been published in the 2024 Special Issue on Computation Resources in the Journal of Molecular Biology: Vallat B et al., IHMCIF: An Extension of the PDBx/mmCIF Data Standard for Integrative Structure Determination Methods, J Mol Biol. 2024; 168546. doi: 10.1016/j.jmb.2024.168546.
April 17, 2024
We are excited to share that we have updated PDB-Dev validation reports to version 1.1. The key feature of this update is the integration of the validation report generation workflow with the deposition and curation pipeline. Validation reports are now generated during the submission process, along with the issue of accession codes, making these reports available for depositors to review and share pre-release. Other changes include:
- Update of MolProbity to version 4.5.2
- Minor changes to the presentation of SAS plots
- Better support for sasCIF format
April 17, 2024
CASP (Critical Assessment of protein Structure Prediction) challenges are held every two years. The CASP organizers are now requesting targets for the upcoming 2024 CASP16 challenge in the following categories.
- Single protein structures
- Protein complexes
- Nucleic acids structures and complexes
- Organic ligand-protein complexes
- Ensembles of macromolecule conformations
- Integrative modeling
CASP is only possible with the generous participation of the experimental structural biology community in providing suitable targets. CASP targets can be directly submitted to CASP organizers or to the PDB. For more information regarding these submissions, please visit the CASP16 website or the wwPDB news site.
We welcome submission of CASP targets belonging to the integrative modeling category to PDB-Dev. Please notify the submission of CASP targets to the PDB-Dev team via email at pdb-dev@mail.wwpdb.org.
January 29, 2024
Starting January 22nd 2024, PDB-Dev accepts automated depositions of integrative structures via an API. Depositors can use a command line interface (CLI) tool to upload IHMCIF files and create new entries for submission. The tool has been implemented using the deriva-py python API provided by the DERIVA scientific asset management platform. The CLI tool is particularly useful for bulk uploads and for creating multiple entries belonging to a collection. Please refer to our deposition documentation for more information.
March 28, 2023
The PDB-Dev team is excited to announce a new feature - structure collections - to better reflect the links between structures generated within the scope of a single research project, investigation, or publication. With increasing numbers of structures generated through high-throughput experimental methods and artificial intelligence-based computational tools, such as AlphaFold and RoseTTAFold, structure collections allow for a convenient way of linking related entries. A set of entries can be identified as belonging to a collection during the deposition process. A collection identifier is issued during biocuration, and a list of associated entries can be obtained at any time by querying the collection identifier.
We thank Andrea Graziadei from the Rappsilber lab for helping us implement and test structure collections. Users are encouraged to explore the first collection of entries in PDB-Dev to learn more about this feature. The accompanying paper describes the development of a novel integrative modeling method using in-cell photo crosslinking mass spectrometry and deep learning.
October 1, 2022
We are proud to announce an important milestone for PDB-Dev - the first 100 entries are released to the public. This achievement is a result of the continued collaboration among the PDB-Dev team, members of the wwPDB IHM task force, wwPDB leadership, RCSB PDB team members, and our pioneer depositors, who were the first to realize the importance of sharing the results of integrative structural biology investigations. The PDB-Dev project is funded by the United States National Science Foundation.
The first 100 entries also offer us a glimpse of the complex landscape of modern integrative modeling. While experimental models used with chemical cross-linking are a leading category, techniques such as three-dimensional electron microscopy (3DEM), small angle solution scattering (SAS), Forster resonance enery transfer (FRET), and others, together with powerful AI-based methods for the de novo structure prediction, give us more than 50(!) unique combinations of datasets used for integrative modeling (see figure below). Because integrative structures provide a unique opportunity to gain insights into how biomolecules function in conditions close to the native, many of these investigations are featured in journals such as Cell, Nature, Science, Proceedings of the National Academy of Sciences, and the Journal of Molecular Biology.
We designed PDB-Dev according to the FAIR guiding principles to make the integrative structures of biomolecules Findable, Accessible, Interoperable, and Reusable, and with the your help, we will continue to improve it and provide better archiving and structure validation services to our users.
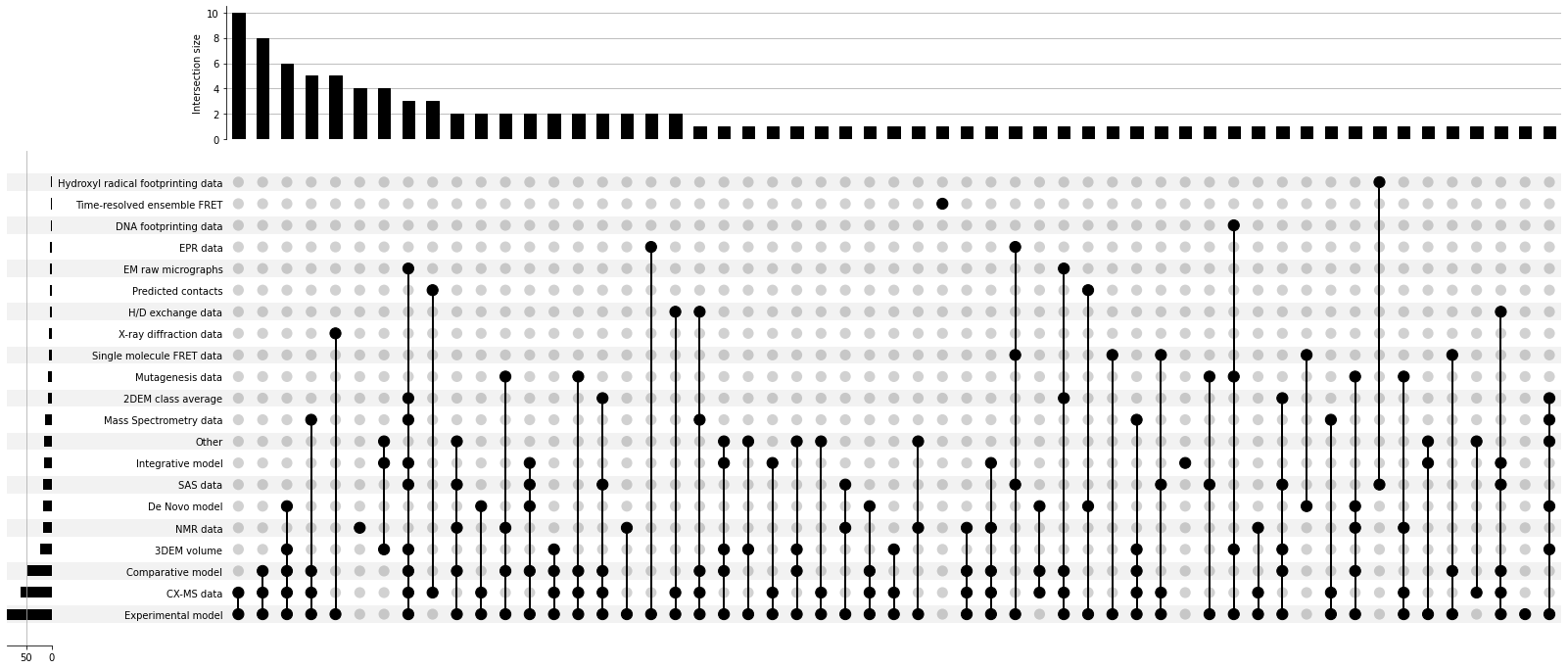

Combinations of datasets used for integrative modeling studies and their frequencies in PDB-Dev entries
September 3, 2022
For a long time, the primary sources of starting structures for integrative modeling were experimental structures from the Protein Data Bank or computational models generated using various comparative modeling approaches. The recent machine learning revolution has led to the development of modeling methods like AlphaFold2 and RoseTTAFold, and also brought a whole new universe of high-quality predicted structures of proteins. More than 200 million predicted structures of individual proteins and complexes are now available to researchers across multiple databases. Now these structures are finally making their way into integrative modeling.
In PDBDEV_00000141, Noone et. al., report a structure of pentraxin protein PTX3, a member of a family of soluble pattern recognition molecules that form an important part of innate immunity, where they facilitate the response to infections and damage by triggering processes such as inflammation. The complete structure of PTX3 was built by integrative modeling using three-dimensional electron microscopy map, mass spectrometry data, and AlphaFold-based starting models.
July 8, 2022
Based on the recommendations of the wwPDB IHM TaskForce (Berman et al. 2019), we have developed validation reports for structures archived in PDB-Dev. The first version of the validation reports have been released on July 8th, 2022. Additional documentation and information about the validation reports are also provided.
The validation report consists of four categories: (i) model composition, (ii) data quality assessments, (iii) model quality assessments, and (iv) fit to data used to build the model. At present, data quality assessments and fit to data used to build the models are evaluated for models built using Small Angle Scattering (SAS) data based on the guidelines published by the wwPDB SAS validation task force.
Validation of integrative structures is a work in progress and additional criteria for validating models based on different experimental methods (such as FRET, chemical crosslinking, 3DEM) will be incorporated in subsequent versions of the validation report.
Please reach out us via email if you have any questions or comments.
July 8, 2022
We have updated the embargo (HOLD) policy for structures deposited to PDB-Dev. Following wwPDB policies, entries can be placed on hold for a maximum period of one year from the date of deposition. If an entry remains unreleased at the end of the hold period, it must either be released or withdrawn. All unreleased entries that have been on HOLD for more than a year have been released by June 24th, 2022. The updated embargo policy is effective from July 1st 2022.
June 18, 2021
We have developed a new Deposition and Data Harvesting System that provides a web interface for depositors to assemble all the information required for archiving integrative structures in PDB-Dev, generate a compliant mmCIF file, and deposit the structure to PDB-Dev. This includes the submission of integrative structures, associated spatial restraints and starting models used, modeling protocols and metadata information (citations, authors, software, data in external repositories, reference sequence information etc.).
The system provides an automated mechanism for collecting diverse and heterogeneous types of integrative structures and restraint data. It has been developed using the DERIVA scientific asset management platform and is based on the definitions in the IHMCIF dictionary and the parent PDBx/mmCIF dictionary.
A User Guide has been created with documentation regarding creating accounts and submitting structures.
Users are encouraged to provide any feedback via email to pdb-dev@mail.wwpdb.org.
The old deposition system that had rudimentary file upload functionality, has been deprecated as of June 3rd, 2022.
December 15, 2021
Recent updates to PDB-Dev, including the development of a new data harvesting system, have been published in Acta Crystallographica Section D: Vallat B et al., New sytem for archiving integrative structures, Acta Cryst. 2021; D77: 1486-1496. doi:10.1107/S2059798321010871
July 2, 2020
Recently, the PDB-Dev team participated in the BioExcel webinar series. The webinar presentation was titled "PDB-Dev: A prototype system for archiving integrative structures". The recorded version of the webinar is available on youtube.
May 20, 2020
3D visualization of structures using Molstar is now available on PDB-Dev. Structures in PDB-Dev can be directly visualized from the respective entry pages. Molstar can visualize atomic as well as multi-scale structures.
Integrative structures can also be visualized online using the Molstar viewer.
Molstar is an open collaboration started by PDBe and RCSB PDB to provide a technology stack for data delivery and analysis tools of macromolecules.
Molstar is under active development. Please report any issues on the Molstar GitHub site.
March 4, 2020
The PDB-Dev web interface has been revamped to provide dynamic, responsive and mobile-friendly web pages. The newly designed website has been made possible due to an overhaul of our backend infrastructure.
The PDB-Dev website now includes a new service that facilitates search and retrieval of integrative structures archived in PDB-Dev. Simple search using macromolecular names (e.g., nuclear pore complex), entry identifiers (e.g., PDBDEV_00000012), experimental methods used (e.g., NMR, 3DEM, Crosslinking-MS), author names (e.g., Sali), software used (e.g., IMP, HADDOCK, ROSETTA), and several other keywords are now supported. Search results can be downloaded as CSV or EXCEL files.
We have also added individual entry pages that provide a summary of the structure (title, description, authors) along with other details regarding the citation, software, input data, and links to related resources and downloading the structure.
We welcome feedback from our users. Please send any comments or suggestions to pdb-dev@mail.wwpdb.org.
March 4, 2020
In March 2019, a satellite workshop titled “Working towards federating structural models and data” was held at the Biophysical Society Annual meeting in Baltimore, Maryland to assess current progress and discuss further requirements for archiving integrative structures. The primary goal of the workshop was to bring together experts from different scientific communities contributing to integrative structural biology and build consensus for addressing the challenges involved in creating common data standards, building methods for federated data exchange, and developing mechanisms for validation of integrative structures. A whitepaper summarizing the outcomes of the workshop was recently published in the journal Structure.
Berman HM et al., Structure. 2019 Dec; 27(12):1745-1759. doi: 10.1016/j.str.2019.11.002.
March 4, 2020
The multi-scale integrative structures downloaded from PDB-Dev can be visualized using the ChimeraX software.
ChimeraX is under active development. It is recommended to use the Daily Build of ChimeraX for visualization.
Please refer to the Daily Build Notes for information regarding operating systems currently supported by ChimeraX.
The integrative structures can be downloaded as text files (use right click + save as) and opened on ChimeraX using the "format ihm" command line option along with the complete path to the downloaded file:
open path/to/myfile.cif format ihm
Alternately, the following command can be used to directly visualize an entry from PDB-Dev, eg: PDBDEV_00000010.
open 10 from pdbdev ignoreCache true
Recent Updates
The PDB-Dev entry files have been recently updated to comply with updates in the IHMCIF dictionary.
Older files downloaded from PDB-Dev will not be supported on the latest ChimeraX. Similarly, the new PDB-Dev files will not be supported on older versions of ChimeraX.
Please download a fresh copy of the file from PDB-Dev and/or clear the ChimeraX cached IHM files (delete ~/Downloads/ChimeraX/PDBDev) for visualizing the updated files with the latest ChimeraX.